My research interests focus on understanding the interactions between proteins and their ligands or substrates, with particular emphasis on understanding the forces that lead to ligand binding, how these forces are used to promote enzymatic catalysis, and how our knowledge of protein-ligand interactions can be used to design more potent inhibitors. My experimental approach involves synthesis of substrates and substrate analogs that can be used in binding and enzymatic studies. My current research projects are described below:
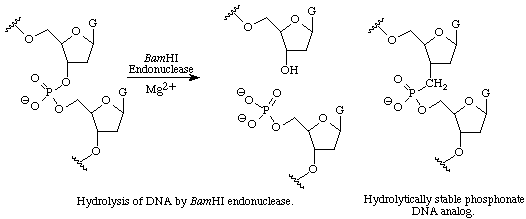
Restriction endonucleases comprise a large class of enzymes that cleave double stranded DNA at specific sites leaving 5' phosphates. Although these enzymes have played a critical role in the development of modern molecular biology, relatively little is known about the molecular mechanism by which they cleave DNA. We are synthesizing phosphonate DNA analogs containing a non-hydrolyzable P-CH2 bond that can be used in crystallographic studies with BamHI or other endonucleases.
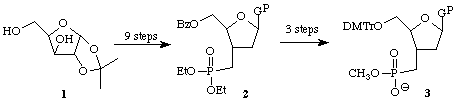
The synthesis of phosphonate nucleotide analogs is quite challenging. We have succeeded in preparing the phosphonate nucleoside analog 2 in several steps from the commercially available sugar 1. Further functional group manipulation will produce the protected nucleotide analog 3 suitable for incorporation into an oligonucleotide using an automated DNA synthesizer.
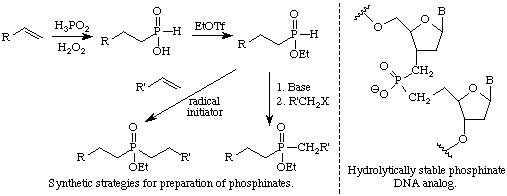
Recent studies show that short strands of synthetic DNA analogs are able to prevent the expression of specific genes. There are a number of studies investigating the use of such oligonucleotides as therapeutic agents. One requirement of these DNA analogs is that they be hydrolytically stable, since foreign DNA is rapidly degraded when it enters a cell. We are developing synthetic methods for preparing "phosphinate DNA" in which the two P-O bonds in the phosphodiester linkage of DNA are replaced with hydrolytically stable P-C bonds. Eventually we hope to develop a procedure for convenient production of short oligonucleotides in which all the P-O bonds in the backbone are replaced by P-C bonds.
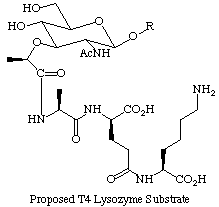
From a structural standpoint, the lysozyme from T4 bacteriophage is one of the most thoroughly studied enzymes. Brian Matthews and coworkers use this enzyme as a system in which to study ideas about protein stability; numerous mutants have been made, analyzed for their stability, and their crystal structures determined at high resolution. Despite this wealth of structural data, many questions about the substrate specificity, kinetics, and mechanism of T4 lysozyme remain unanswered, due in large part to the complexity of the substrate and the lack of a convenient small molecule assay. We are synthesizing small oligosaccharide fragments of the cell wall to probe the kinetics and substrate specificity of T4 lysozyme. The ability to synthesize substrates of T4 lysozyme will allow us to develop a convenient, continuous fluorescence assay that will facilitate kinetic studies on various mutant enzymes. Eventually we hope to synthesize non-hydrolyzable substrate analogs to probe the mechanism of this enzyme.